- Exon count:
- 6
| Annotation release |
Status |
Assembly |
Chr
|
Location |
| RS_2023_10 |
current |
GRCh38.p14 (GCF_000001405.40) |
16 |
NC_000016.10 (366969..370538, complement) |
| RS_2023_10 |
current |
T2T-CHM13v2.0 (GCF_009914755.1) |
16 |
NC_060940.1 (361841..366186, complement) |
| 105.20220307 |
previous assembly |
GRCh37.p13 (GCF_000001405.25) |
16 |
NC_000016.9 (416969..420538, complement) |
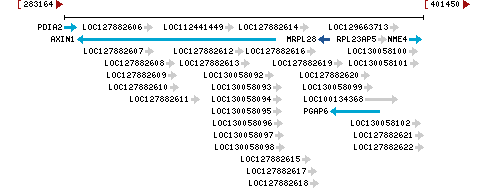
Chromosome 16 - NC_000016.10